RNSGA-II Searcher example
This is an example about how using RNSGAIISearcher for multi-objectives optimization.
1. Import modules and prepare data
[1]:
from hypernets.core.random_state import set_random_state
set_random_state(1234)
from hypernets.utils import logging as hyn_logging
from hypernets.examples.plain_model import PlainModel, PlainSearchSpace
from hypergbm import make_experiment
from hypernets.tabular import get_tool_box
from hypernets.tabular.datasets import dsutils
from hypernets.tabular.sklearn_ex import MultiLabelEncoder
#hyn_logging.set_level(hyn_logging.WARN)
df = dsutils.load_bank().head(1000)
tb = get_tool_box(df)
df_train, df_test = tb.train_test_split(df, test_size=0.2, random_state=9527)
2. Run an experiment within NSGAIISearcher
[2]:
import numpy as np
experiment = make_experiment(df_train,
eval_data=df_test.copy(),
callbacks=[],
random_state=1234,
search_callbacks=[],
target='y',
searcher='rnsga2', # available MOO searchers: moead, nsga2, rnsga2
searcher_options=dict(ref_point=np.array([0.1, 2 ]), weights=np.array([0.1, 2]), population_size=10),
reward_metric='logloss',
objectives=['nf'],
early_stopping_rounds=30,
drift_detection=False)
estimators = experiment.run(max_trials=30)
hyper_model = experiment.hyper_model_
hyper_model.searcher
[2]:
RNSGAIISearcher(objectives=[PredictionObjective(name=logloss, scorer=make_scorer(log_loss, needs_proba=True), direction=min), NumOfFeatures(name=nf, sample_size=1000, direction=min)], recombination=SinglePointCrossOver(random_state=RandomState(MT19937))), mutation=SinglePointMutation(random_state=RandomState(MT19937), proba=0.7)), survival=_RDominanceSurvival(ref_point=[0.1 2. ], weights=[0.1 2. ], threshold=0.3, random_state=RandomState(MT19937))), random_state=RandomState(MT19937)
3. Summary trails
[3]:
df_trials = hyper_model.history.to_df().copy().drop(['scores', 'reward'], axis=1)
df_trials[df_trials['non_dominated'] == True]
[3]:
| trial_no | succeeded | elapsed | non_dominated | model_index | reward_logloss | reward_nf | |
|---|---|---|---|---|---|---|---|
| 4 | 5 | True | 0.386380 | True | 0.0 | 0.217409 | 0.625 |
| 9 | 10 | True | 0.798336 | True | 1.0 | 0.218916 | 0.4375 |
| 11 | 12 | True | 0.682108 | True | 2.0 | 0.287458 | 0.0 |
| 14 | 15 | True | 0.455569 | True | 3.0 | 0.221304 | 0.375 |
| 26 | 28 | True | 0.568367 | True | 4.0 | 0.045741 | 0.9375 |
| 28 | 30 | True | 0.407018 | True | 5.0 | 0.141022 | 0.6875 |
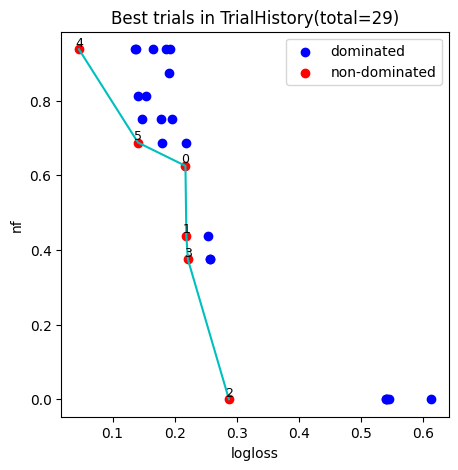
4. Plot pareto font
We can pick model accord to Decision Maker’s preferences from the pareto plot, the number in the figure indicates the index of pipeline models.
[4]:
fig, ax = hyper_model.history.plot_best_trials()
fig.show()
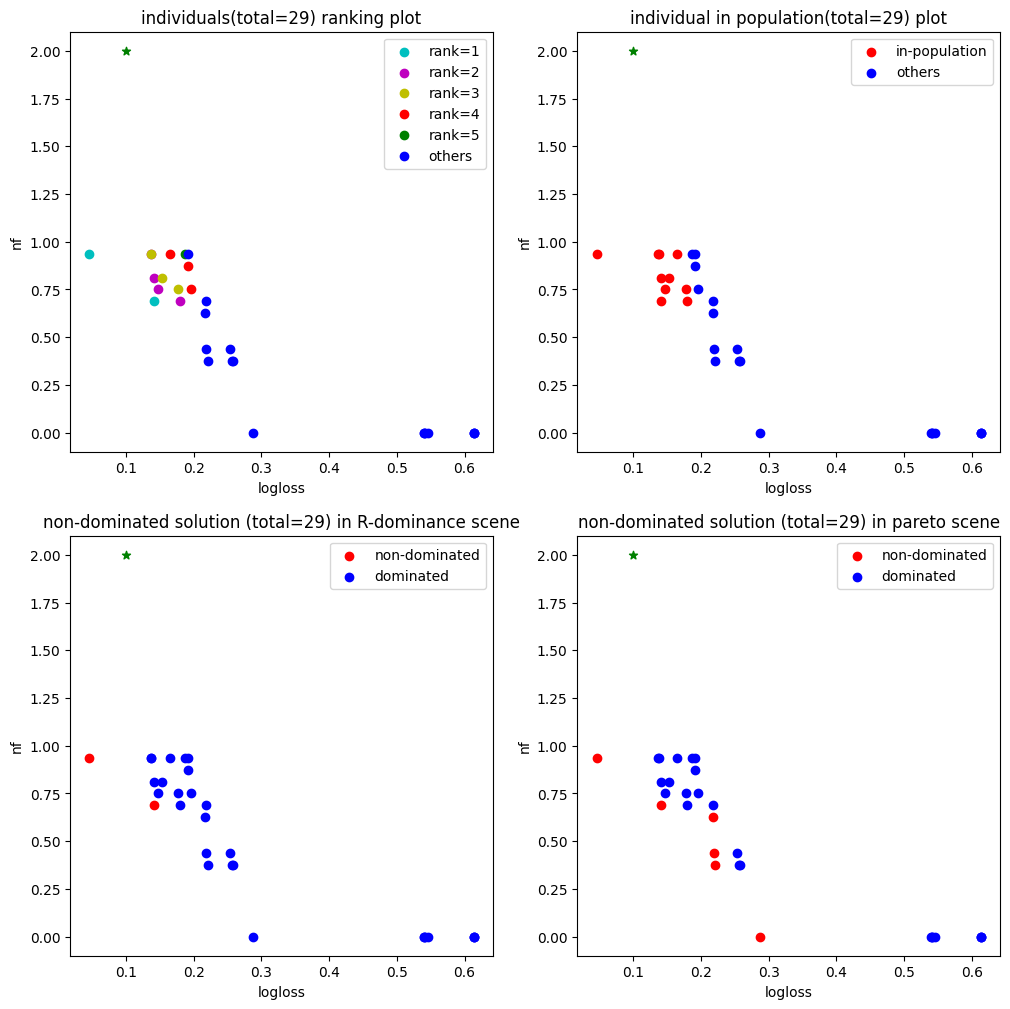
5. Plot population
[5]:
fig, ax = hyper_model.searcher.plot_population()
fig.show()
6. Evaluate the selected model
[6]:
print(f"Number of pipeline: {len(estimators)} ")
pipeline_model = estimators[0] # selection the first pipeline model
X_test = df_test.copy()
y_test = X_test.pop('y')
preds = pipeline_model.predict(X_test)
proba = pipeline_model.predict_proba(X_test)
tb.metrics.calc_score(y_test, preds, proba, metrics=['auc', 'accuracy', 'f1', 'recall', 'precision'], pos_label="yes")
Number of pipeline: 6
[6]:
{'auc': 0.8417038690476191,
'accuracy': 0.855,
'f1': 0.17142857142857143,
'recall': 0.09375,
'precision': 1.0}